This web page was produced as an assignment for Genetics 564 at UW-Madison in Spring 2017.
Introduction
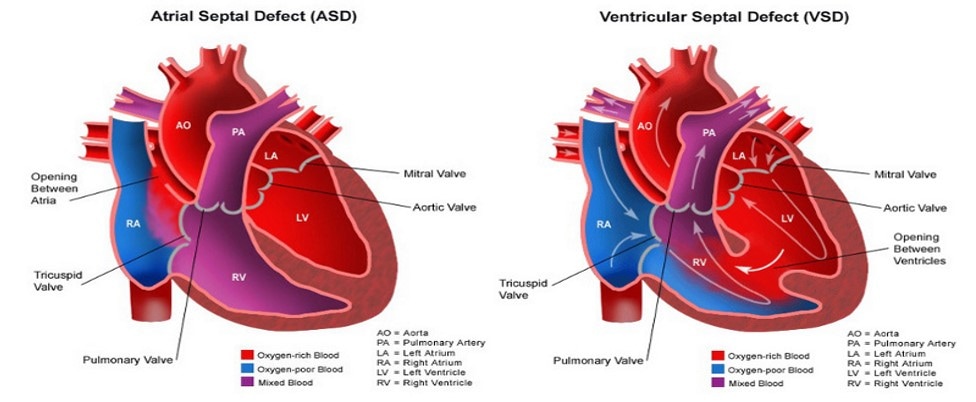
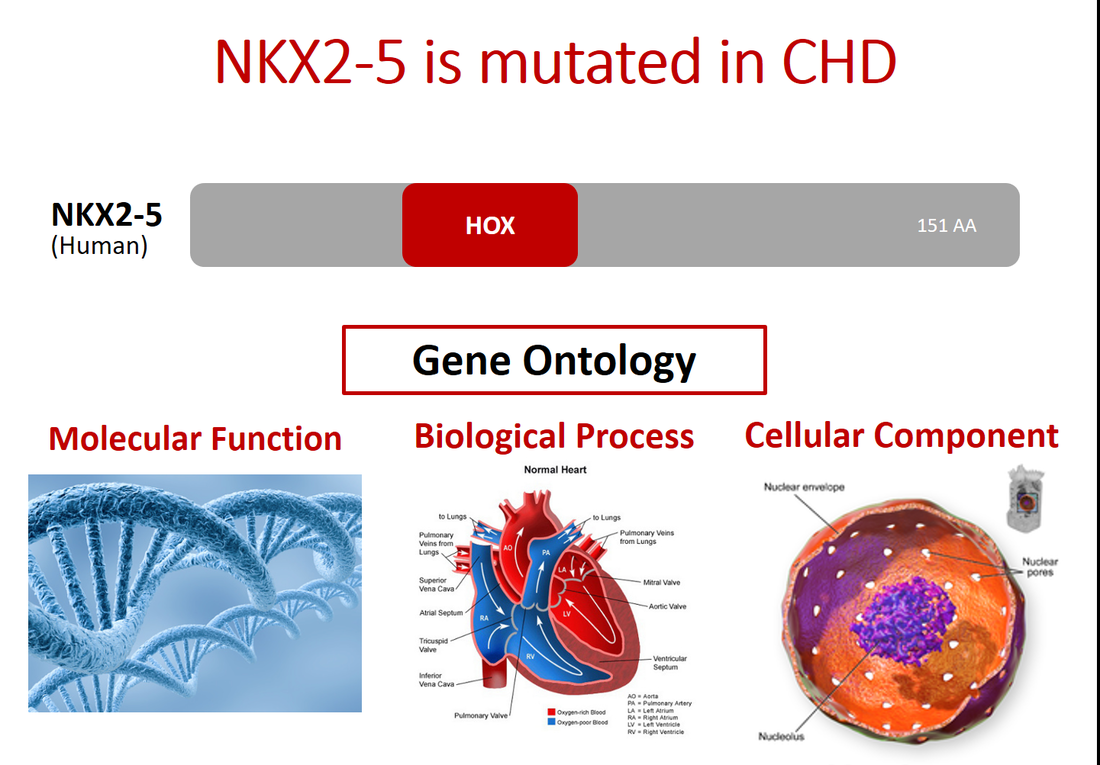
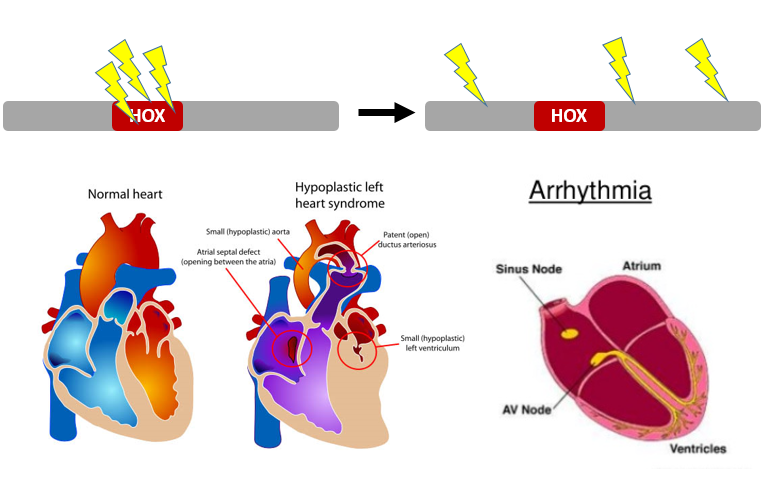
Congenital heart disease (CHD) accounts for the largest percentage of morbid congenital abnormalities in the world, affecting approximately 8 out of every 1000 births [1]. CHD is caused by the disruption of a complex network of cardiac regulatory genes which dictate proper heart formation [2]. The NKX2-5 gene, a master regulator of heart formation in vertebrates, encodes a homeobox transcription factor that interacts with several cardiac regulatory genes [3]. Previous studies have shown that specific mutations in NKX2-5 lead to characteristic defects of CHD in humans, such as septal defects, arrhythmias, or eventual heart failure [4]. Despite this detailed knowledge of how specific mutations in NKX2-5 give rise to CHD defects, the role of NKX2-5 in cardiac morphogenesis remains poorly understood.
My primary goal is to determine the mechanism by which mutations in the NKX2-5 HOX region (homeodomain) give rise to septal defects. Mice have highly similar genetic and physiological characteristics to those of humans, a short life span, and a high breeding rate [5]. For these primary reasons, mice will be used as the organism to model CHD found in humans. I hypothesize that mutations in the NKX2-5 homeodomain disrupt downstream cardiac genes and its protein-protein interaction network, leading to septal defects of CHD. My long-term goal is to uncover the function of the entire NKX2-5 gene in septal defects and proper heart formation.
Specific Aims
Aim 1
Aim 1: Determine if NKX2-5 HOX mutations directly cause septal defects.
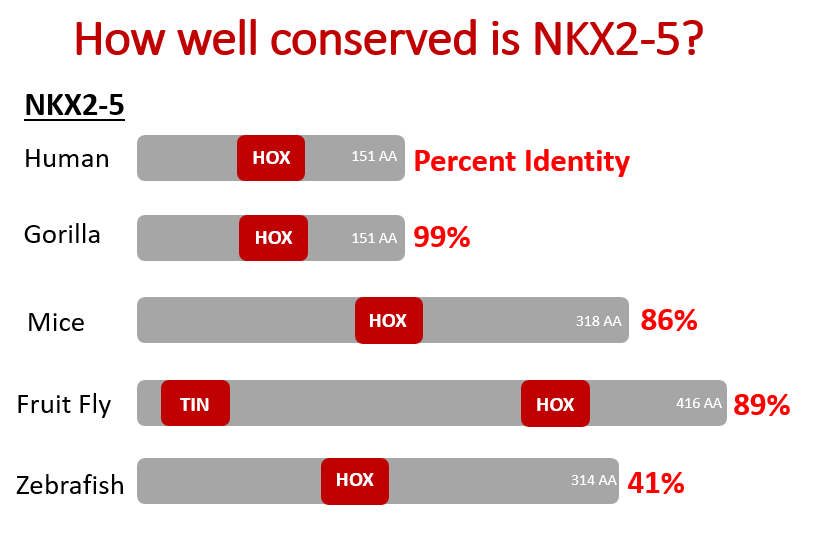
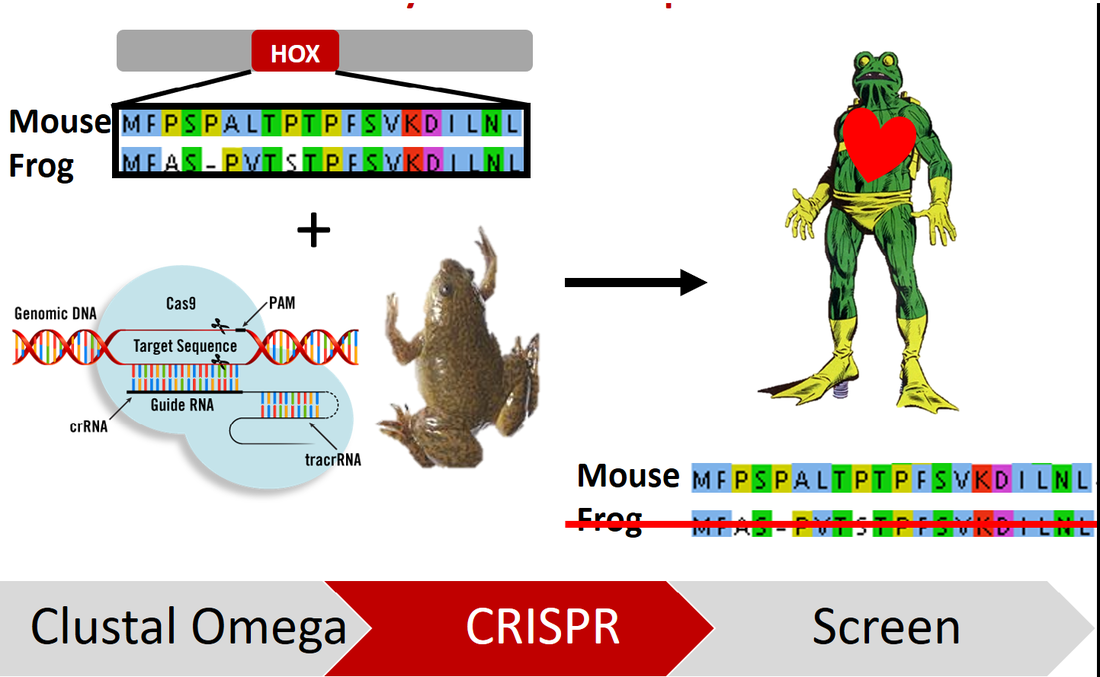
Approach: ClustalOmega and BLAST will be used to identify conserved regions in the NKX2-5 homeodomain between model organisms with three- and four-chambered hearts. Then, the homeodomain sequence from a four-chambered heart organism will be inserted into the NKX2-5 homeodomain of a three-chambered heart organism using CRISPR/Cas9. Mutations that have been found in humans with septal defects will then be induced in the modified organism.
Hypothesis: I hypothesize that mutations in the NKX2-5 homeodomain found in four-chambered hearts is directly responsible for the formation of septal defects.
Rationale: The NKX2-5 homeodomain is highly conserved in four-chambered heart organisms and three-chambered heart organisms are unable to form ventricular septal defects.
Aim 1: Determine if NKX2-5 HOX mutations directly cause septal defects.
Approach: ClustalOmega and BLAST will be used to identify conserved regions in the NKX2-5 homeodomain between model organisms with three- and four-chambered hearts. Then, the homeodomain sequence from a four-chambered heart organism will be inserted into the NKX2-5 homeodomain of a three-chambered heart organism using CRISPR/Cas9. Mutations that have been found in humans with septal defects will then be induced in the modified organism.
Hypothesis: I hypothesize that mutations in the NKX2-5 homeodomain found in four-chambered hearts is directly responsible for the formation of septal defects.
Rationale: The NKX2-5 homeodomain is highly conserved in four-chambered heart organisms and three-chambered heart organisms are unable to form ventricular septal defects.
Aim 2
Aim 2: Determine the role of NKX2-5 homeodomain mutations on the formation of septal defects and the gene expression of cardiac regulatory proteins.
Approach: Homeodomain mutations that are known to cause septal defects in humans will be induced in mice models. The mice that are found to have septal defects will have their heart tissue sampled and analyzed with RNA-seq.
Hypothesis: There will be significant changes in the transcription levels of NKX2-5 and its interacting cardiac regulatory proteins.
Rationale: NKX2-5 both positively and negatively regulates numerous genes during heart formation. Mutations in NKX2-5 can result in ectopic levels of downstream interacting proteins, leading to CHD.
Aim 2: Determine the role of NKX2-5 homeodomain mutations on the formation of septal defects and the gene expression of cardiac regulatory proteins.
Approach: Homeodomain mutations that are known to cause septal defects in humans will be induced in mice models. The mice that are found to have septal defects will have their heart tissue sampled and analyzed with RNA-seq.
Hypothesis: There will be significant changes in the transcription levels of NKX2-5 and its interacting cardiac regulatory proteins.
Rationale: NKX2-5 both positively and negatively regulates numerous genes during heart formation. Mutations in NKX2-5 can result in ectopic levels of downstream interacting proteins, leading to CHD.
Aim 3
Aim 3: Identify NKX2-5 homeodomain mutations that cause septal defects and disrupt protein-protein interactions of NKX2-5.
Approach: STRING will be used to identify protein interactors of NKX2-5. CRISPR/Cas9 will then be used to induce known mutations that cause septal defects in mice. Mice that develop septal defects will have their heart tissue sampled for tandem mass spectrometry and western blot.
Hypothesis: The homeodomain mutations will result in a decrease in the interaction of NKX2-5 with known protein-protein interactors.
Rationale: The conserved amino acids in NKX2-5 are essential for NKX2-5 to bind to or be bound by known interacting proteins.
Aim 3: Identify NKX2-5 homeodomain mutations that cause septal defects and disrupt protein-protein interactions of NKX2-5.
Approach: STRING will be used to identify protein interactors of NKX2-5. CRISPR/Cas9 will then be used to induce known mutations that cause septal defects in mice. Mice that develop septal defects will have their heart tissue sampled for tandem mass spectrometry and western blot.
Hypothesis: The homeodomain mutations will result in a decrease in the interaction of NKX2-5 with known protein-protein interactors.
Rationale: The conserved amino acids in NKX2-5 are essential for NKX2-5 to bind to or be bound by known interacting proteins.
Future Directions
References
Images
1.https://advancedcardioservices.com/wp-content/uploads/2016/05/anti-arrhythmic-drug-thereapy-12-638.jpg
2.http://qsota.com/wp-content/uploads/2015/03/Congenital-heart-disease-causes-symptoms-and-treatment.jpg
3.http://media.istockphoto.com/vectors/pile-of-used-papers-vector-id635477438
4.https://d7umqicpi7263.cloudfront.net/img/product/0c8ac6ea-9c49-426e-894d-c299a53b4914/9660d7e5-88bb-4009-bc27-49e83ed20a03.jpg
5.https://www.mirusbio.com/assets/figures/crispr-cas9-genome-editing.png
6.http://www.writeups.org/wp-content/uploads/Frog-Man-Marvel-Comics-Spider-Man-a.jpg
7.https://www.synthego.com/wp-content/uploads/2016/07/crispr-cas9.png
8.http://news.nationalgeographic.com/news/2006/07/images/060705-mouse-frog_big.jpg
Content
1.http://atlasgeneticsoncology.org/Genes/NKX25ID42958ch5q35.html
2.https://blast.ncbi.nlm.nih.gov/Blast.cgi
3.http://www.ideacenter.org/contentmgr/showdetails.php/id/1113
4.http://www.ebi.ac.uk/Tools/services/web/toolresult.ebi?jobId=clustalo-I20170427-044430-0152-63375539-oy&analysis=summary
5.Benson, D. Woodrow, et al. "Mutations in the cardiac transcription factor NKX2. 5 affect diverse cardiac developmental pathways." The Journal of clinical investigation 104.11 (1999): 1567-1573.
6.Chung, Ill-Min, and Govindasamy Rajakumar. "Genetics of congenital heart defects: The NKX2-5 gene, a key player." Genes 7.2 (2016): 6.
7.Brown, Carl O., et al. "The cardiac determination factor, Nkx2-5, is activated by mutual cofactors GATA-4 and Smad1/4 via a novel upstream enhancer." Journal of Biological Chemistry 279.11 (2004): 10659-10669.
1.https://advancedcardioservices.com/wp-content/uploads/2016/05/anti-arrhythmic-drug-thereapy-12-638.jpg
2.http://qsota.com/wp-content/uploads/2015/03/Congenital-heart-disease-causes-symptoms-and-treatment.jpg
3.http://media.istockphoto.com/vectors/pile-of-used-papers-vector-id635477438
4.https://d7umqicpi7263.cloudfront.net/img/product/0c8ac6ea-9c49-426e-894d-c299a53b4914/9660d7e5-88bb-4009-bc27-49e83ed20a03.jpg
5.https://www.mirusbio.com/assets/figures/crispr-cas9-genome-editing.png
6.http://www.writeups.org/wp-content/uploads/Frog-Man-Marvel-Comics-Spider-Man-a.jpg
7.https://www.synthego.com/wp-content/uploads/2016/07/crispr-cas9.png
8.http://news.nationalgeographic.com/news/2006/07/images/060705-mouse-frog_big.jpg
Content
1.http://atlasgeneticsoncology.org/Genes/NKX25ID42958ch5q35.html
2.https://blast.ncbi.nlm.nih.gov/Blast.cgi
3.http://www.ideacenter.org/contentmgr/showdetails.php/id/1113
4.http://www.ebi.ac.uk/Tools/services/web/toolresult.ebi?jobId=clustalo-I20170427-044430-0152-63375539-oy&analysis=summary
5.Benson, D. Woodrow, et al. "Mutations in the cardiac transcription factor NKX2. 5 affect diverse cardiac developmental pathways." The Journal of clinical investigation 104.11 (1999): 1567-1573.
6.Chung, Ill-Min, and Govindasamy Rajakumar. "Genetics of congenital heart defects: The NKX2-5 gene, a key player." Genes 7.2 (2016): 6.
7.Brown, Carl O., et al. "The cardiac determination factor, Nkx2-5, is activated by mutual cofactors GATA-4 and Smad1/4 via a novel upstream enhancer." Journal of Biological Chemistry 279.11 (2004): 10659-10669.