What is protein phylogeny?
Phylogeny is the evolution of a genetically related group of organisms as distinguished from the development of the individual organism through changes in DNA or protein (amino acid) sequences [1]. To illustrate phylogenetic relationships between organisms, phylogenetic trees are created based on either DNA or amino acid sequence. The overlap of sequences across species or organisms can determine how related or unrelated they are from each other; thus, giving insight into their phylogeny. Below are 4 different phylogenetic trees that compare the protein phylogeny of organisms for the NKX-2.5 amino acid sequence also shown directly below.
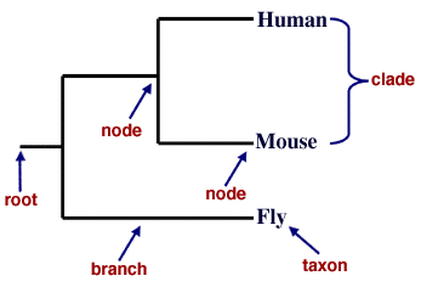
Phylogenetic Tree Basics
Phylogenetic Tree Building
Phylogenetic trees can be constructed in different ways. The following trees for the NKX-2.5 protein were built using the techniques described below.
Percent Identity
This method compares how identical aligned amino acid sequences are across multiple taxa.
BLOSUM Matrix
This method compares how identical sequences are after alignment by utilizing a scored-based system. Each amino acid in a sequence is given a score based on the likelihood of a matching sequence occurring in a random sequence [3].
Average Distance
This method uses equal length branches under the assumption that all taxon diverged similarly from the common ancestor (root).
Neighbor Joining
This method generates actual branch lengths between taxa unlike average distance. It does not assume that all taxon diverged similarly from the common ancestor (root).
Percent Identity
This method compares how identical aligned amino acid sequences are across multiple taxa.
BLOSUM Matrix
This method compares how identical sequences are after alignment by utilizing a scored-based system. Each amino acid in a sequence is given a score based on the likelihood of a matching sequence occurring in a random sequence [3].
Average Distance
This method uses equal length branches under the assumption that all taxon diverged similarly from the common ancestor (root).
Neighbor Joining
This method generates actual branch lengths between taxa unlike average distance. It does not assume that all taxon diverged similarly from the common ancestor (root).
NKX-2.5 Phylogenetic Trees
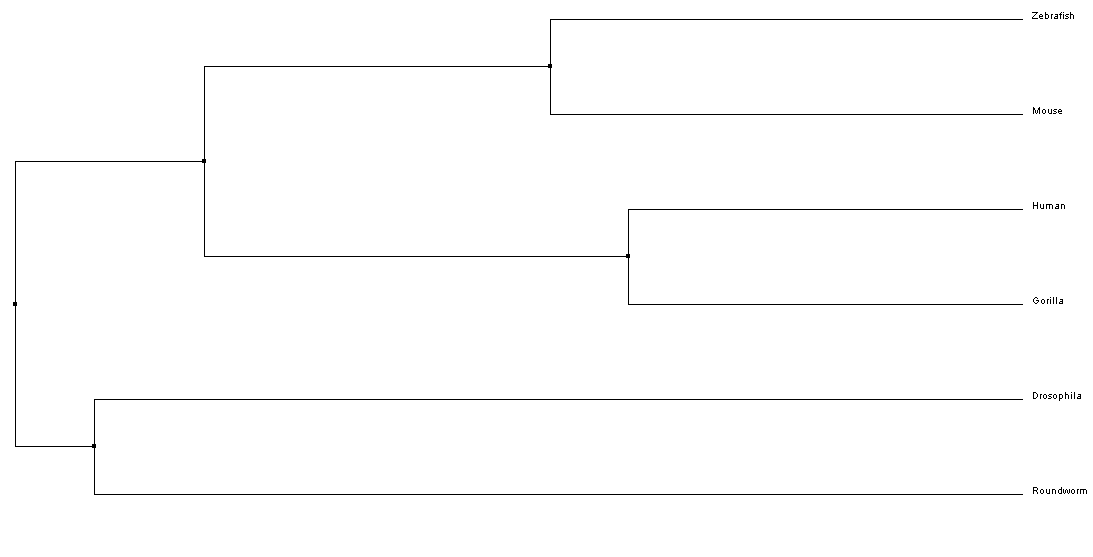
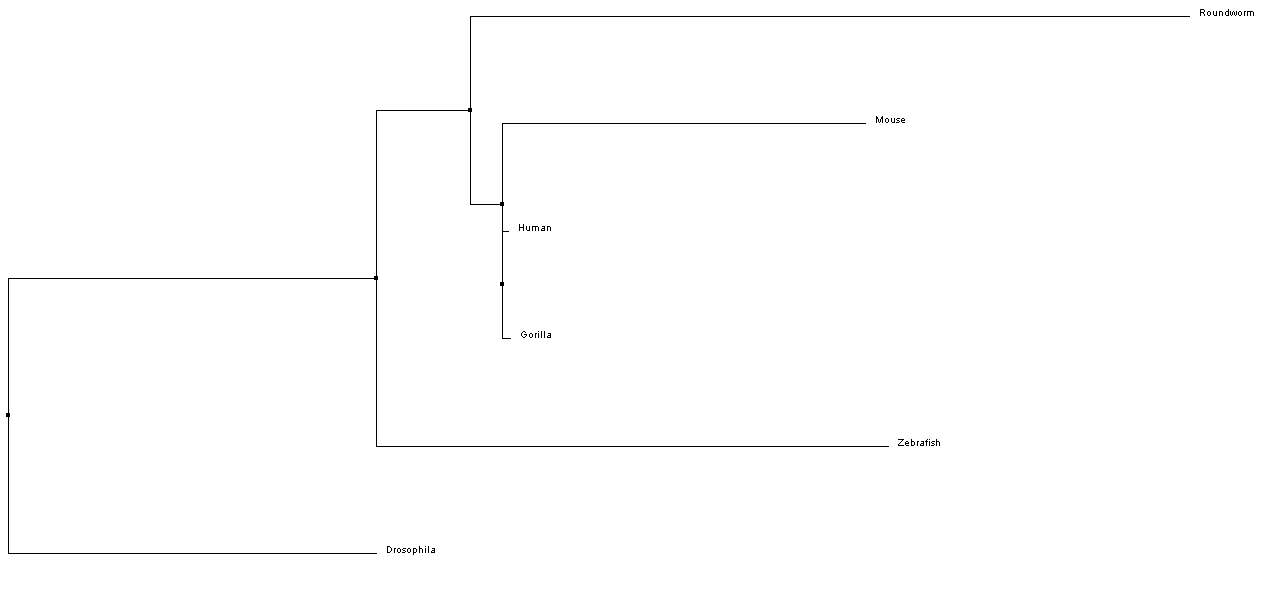
Above is a phylogeny tree of the NKX2-5 protein found in humans and other species. This tree was created with the BLOSUM62/Neighbor Joining method using CustalWOmega and Jalview.
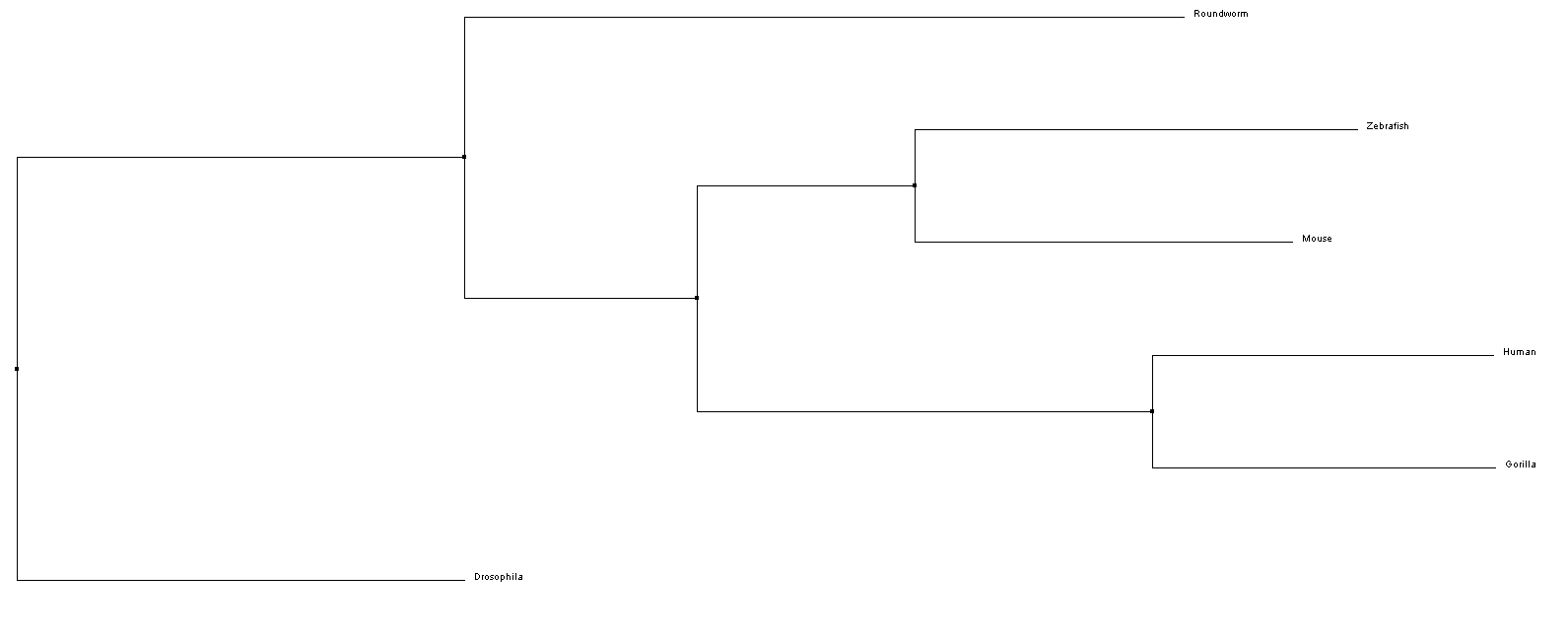
Above is a phylogeny tree of the NKX2-5 protein found in humans and other species. This tree was created with the BLOSUM62/Average Distance method using CustalWOmega and Jalview.
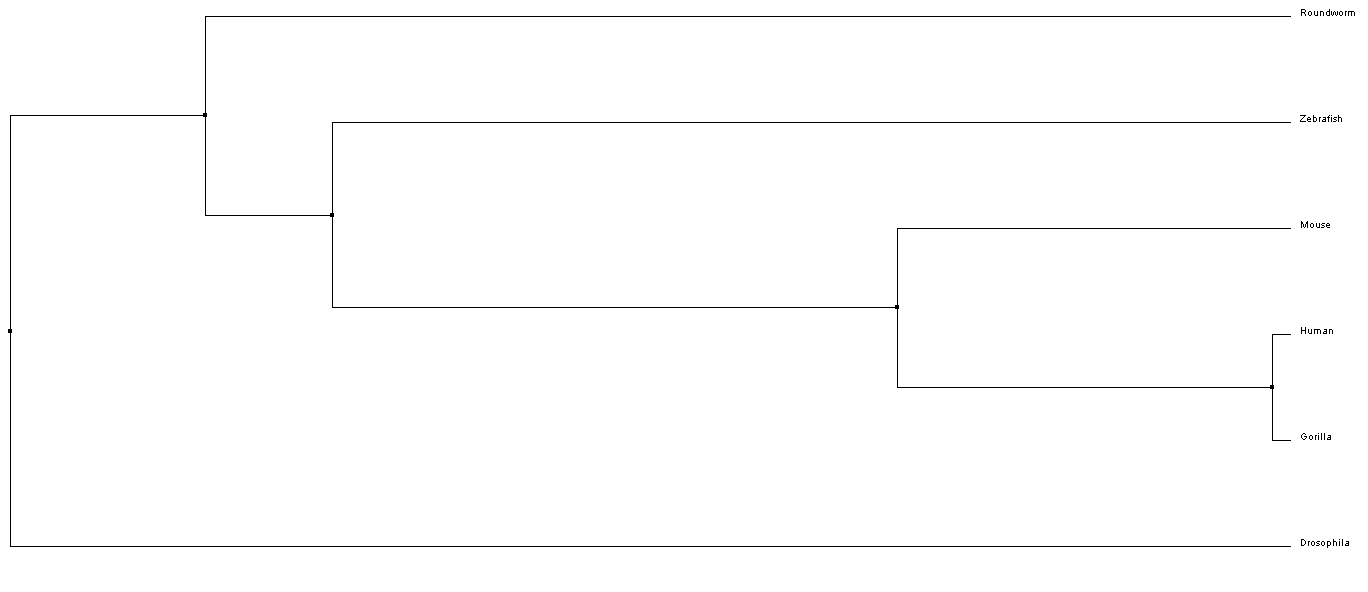
Above is a phylogeny tree of the NKX2-5 protein found in humans and other species. This tree was created with the Percent Identity/Average Distance method using CustalWOmega and Jalview.
Above is a phylogeny tree of the NKX2-5 protein found in humans and other species. This tree was created with the Percent Identity/Neighbor Joining method using CustalWOmega and Jalview.
Conclusions
From the phylogenetic trees of NKX-2.5 in 6 different model organisms, we can conclude that there is the greatest similarity in amino sequence for the NKX-2.5 protein across mammals (i.e. Mice, Humans, and Gorillas). Interestingly, Drosophila is shown to have the least amount of phylogenetic ancestry between taxa. This may suggest that the homolog protein in Drosophila for NKX-2.5 (tinman) functions differently than other species and therefore, has a different amino acid sequence [4].
References
[1] Phylogeny. 2011. In Merriam-Webster.com.Retrieved Feb 14, 2017, from https://www.merriam-webster.com/dictionary/phylogeny.
[2]Skop, A. (n.d.). Homology &Phylogeny. Retrived Feb 14, 2017, from http://genetics564.weebly.com/homology--phylogeny.html
[3] Eddy, S.R.. 2004. Where did the BLOSUM62 alignment score matrix come from? Nature Biotechnology, 1035-1036.
[4] Bodmer, R. (1993). The gene tinman is required for specification of the heart and visceral muscles in
Drosophila. Development, 118(3), 719-729.
[2]Skop, A. (n.d.). Homology &Phylogeny. Retrived Feb 14, 2017, from http://genetics564.weebly.com/homology--phylogeny.html
[3] Eddy, S.R.. 2004. Where did the BLOSUM62 alignment score matrix come from? Nature Biotechnology, 1035-1036.
[4] Bodmer, R. (1993). The gene tinman is required for specification of the heart and visceral muscles in
Drosophila. Development, 118(3), 719-729.